Remote
Sensing Exercise 5
PREPROCESSING
Feature
Extraction and Principal Component Analysis
*Note: IDRISI
commands are specified as follows:
Commands: COMMAND
1 – COMMAND 2 – ETC.
Preprocessing
encompasses a variety of operations designed to make a set of image bands more
manageable, informative and accurate. Processes usually fall into one of the
following categories:
·
Creating
Image Subsets
·
Feature
Extraction
·
Radiometric
Correction
·
Geometric
Correction
Each
of these steps is well documented in Campbell’s chapter 10; Jensen chapters 6
and 7. The purpose of this exercise is to introduce two aspects of preprocessing
- creating “subsets” and “feature extraction” - and to apply these
techniques to the 1986 LANDSAT 5 MSS (Multi-Spectral Scanner) image of
Mexico’s Sierra Basin, a volcanic plateau about 150km southeast of
Guadalajara.
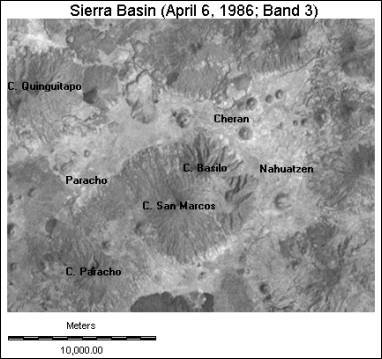
Figure 1:
The study site.
Step
1 - Creating Image Subsets (Campbell pages 290 - 292)
The
study area you have been assigned is depicted in Figure 1. This region of basin contains the towns of Paracho, Cheran
and Nahuatzen, as well as the surrounding volcanic peaks, the largest of which
is Cerro de Paracho. Your objective is to develop a land cover inventory of the
area, including forested land, urban areas and agricultural regions.
The
original LANDSAT image covers a much greater spatial extent than the study area.
By reducing the image such that it contains only the data regarding the
specified area – create a “subset” – the complexity of analysis will be
reduced, storage requirements and processing times will be limited, and results
will likely be easier to interpret and verify.
Before proceeding, make sure that you have copies of the
original MSS bands in your Lab #5 directory (each image has a .rst and .rdc
file):
Band 1 (0.5
- 0.6 micrometers, visible green )
Band 3 (0.7
- 0.8 micrometers, near infrared )
Band 4 (0.8
- 1.1 micrometers, near infrared )
Display
the Band 3 image with the Grey Scale palette with the “Autoscale” option and
no “Title”. Compare the extent of this image to the study area pictured in Figure
1.
The original
image data used in this exercise is actually a subset of an April 20, 1986,
LANDSAT 5 MSS image. The full image represents a 185 X 185-kilometer swath –
roughly 35,000 square kilometers in area. (The original 90-meter resolution has
also been re-sampled to 60-meters for topographic correction and registration
purposes.)
Step
1: Creating image subsets
1.
Set the Project
Environment to the directory containing the Lab 5 folder.
2.
Commands:
REFORMAT – WINDOW
3.
Set
number of files to 4.
4.
Select
the each of the four input images by clicking the browse icon.
5.
In
“Output prefix” Select an appropriate prefix ( refor ) for output files to
create a unique identifier. Click
on “add prefix to file name”
6.
In
“Window specified by” select geographical positions option.
This option allows the user to enter minimum/maximum UTM coordinates (in
meters) of the subset image.
7.
Enter the
UTM positions, as specified in the Table
1, in the appropriate box.
8.
Leave
other settings as default.
9.
Select OK
to process the images.
TABLE 1: Subset Image Coordinates
|
|
Minimum |
Maximum |
X |
174,995
m |
200,010
m |
|
Y |
2,165,990
m |
2,186,025
m |
The
result should be a set of four images - MSS Bands 1 through 4 - that now contain
only the area selected as the study site. Display the new Band 3 image using the
Grey Scale palette and “Autoscale.” It should look similar to Figure
1.
Take
a few minutes and zoom in on Figure 1 to find the area that is located.
Questions:
1)
The
METADATA command [Commands: FILE -
METADATA] provides information on the projection, file structure, size, and
resolution of an image. Compare the metadata of the original image to that of
the subset image - what has changed?
2)
How many
pixels make up the original image and the subset image? Show your work.
3)
What is
the approximate area of the subset image, to the nearest square kilometer? Show
your work.
4)
Display
the Band 3 subset image. Examine the image’s histogram. Why does
“autoscale” produce a good result?
Step
2:
Feature Extraction (Principal Components Analysis) (Campbell pages 287 –
291; Jensen pages 172-179)
When
working with large multi-band imagery data it is often necessary to reduce the
volume of data to only that which is deemed essential for image analysis.
Frequently, different bands of multi-image data contain similar information. Figure
2 illustrates this point. In a highly vegetated area, for example, the use
of both a blue visible and red visible band would most likely be redundant.
Vegetation responds similarly in both of these wavelengths. By removing one of
these bands, the volume of image data is reduced dramatically, yet the amount of
unique information lost may be statistically negligible.
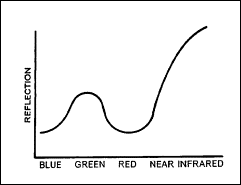
Figure 2: Typical spectral reflectance of vegetation
In
a previous exercise, we examined which bands were highly correlated and
eliminated those redundant ones. If
you remember, however, the correlations between the bands was not perfect and we
perhaps lost some information. Is
there a better way is reduce the data volume (number of bands) we have to
examine, yet maintain the greatest amount of information.
Principal
Component Analysis (PCA) is a mathematical process that can be used to transform
a set of image bands into a new set of virtual bands (called components)
where correlation between the components is minimized. Therefore, each component
contains information different from the others. Each component is a statistical
description of the variability present in all bands, and they are ordered in
terms of the amount of variability they explain. “Component 1” represents
the optimum combination of bands and accounts for the greatest variability in
brightness in the image, “Component 2” the second-most, and so on. This new
data can then be correlated with the original image data to identify the
combination of original bands that contain most of the unique information.
What
is the relation between the spectral bands and the components?
The answer is determined from the “loading”, or how each component
correlates with each band. From a table of load values one can determine the relative
influence of each band on each component. Jensen
includes a good description of this.
Although
this exercise uses a relatively small data set, this is an important and often
used technique in analysis of remotely sensed information. Therefore, we will
perform a basic PCA on the study area image created in Step 1 to determine which original bands contain the most
information. [N.B.
All subsequent exercises will use the study-area images (subset images)
created in Step 1.] The
results will be used in future classification exercises.
1.
Set the Project Environment to
the directory containing the Lab 5 folder.
2.
Command: ANALYSIS - IMAGE
PROCESSING - TRANSFORMATION – PCA
3.
Select “Calculate covariances directly” option.
4.
Set number of files to 4.
5.
Add the reformated image bands by clicking the browse icon for each line.
6.
Select 4 components to be extracted.
It is general practice to extract as many components as there are bands.
7.
Enter a prefix, pca, for the output data
8.
Select the “Unstandardized variables” option.
[A discussion of covariance calculation and
the difference between using unstandardized/standardized variables is beyond the
scope of this exercise. The default settings used by IDRISI are the most
commonly used approach. ]
9.
Click OK to process.
The
results will automatically appear on the screen. The structure of this data is
explained as follows:
VAR/COV -
The variance/covariance matrix. This information is used by the PCA
algorithm to construct the components
COR MATRX -
The correlation matrix. This is the correlation between bands. Bands that
are highly correlated represent much of the same information (a value of 1
represents perfect correlation, a value of 0 no correlation.)
COMPONENT - This is the component summary table. The Eigenvalues express the
amount of total variance in all four image bands explained by each component;
these values are also expressed as a percent of total variance (% VAR.). (The
Eigenvectors are used by the transformation equation; for the purposes of
this exercise, they can be ignored.).
LOADING
- The loadings matrix. The Loadings refers to the degree of correlation
between the original bands with the components. The band correlating most highly
with (and therefore most similar to) component 1 contains the most information
relative to the other original bands. The band correlating most highly with
component 2 is the second-most informative, and so on.
Save
a copy of this table as a textfile; it will be useful in later exercises.
Questions:
Refer to the Correlation Matrix:
10)
Why is
the diagonal equal to 1.0?
11)
Would you
use both Bands 1 and 2 in subsequent analysis? Why or why not? (Describe the
relationship between these bands.)
12)
Explain
one scenario in which your answer to #11 might be different? (Hint: Think about
the spectral response of surface features to the four MSS wavelengths.)
Refer
to the Component Matrix:
13) Examine the “% var.” (percent variance) for each of the components. What do they mean?
14)
How much
of the variance is explained by the first two components?
Show your work.
15)
How much
variance do the final two components explain? Show your work.
At this point, it might be useful to display the 4
PCA component images using the Grey Scale palette and autoscale
16)
For
further analysis, what components would you use and which would you discard?
Explain your answers. Refer
to the histograms for each component image to help you answer this question.
What does Component 4 most likely represent?
17)
What
percentage of data would you be discarding?
Refer
to the Loadings Matrix:
18)
What
original band image(s) most resembles Component 1? Discuss.
19)
What
original band image(s) most resembles Component 2? Discuss.
20)
What
original band image(s) most resembles Component 3 and 4? Discuss
21)
Which
three original bands contain the most information regarding the study site?
Given their correlation, and the loading, did you learn anything from the PCA
analysis that you otherwise might not have done if you were going to select the
manually for analysis?
Refer
to LAB #4:
22)
Assume
that you can only use two of the three SPOT image bands (pdxspotm) for image
classification. Which bands would you use?
23)
How much
information would you be retaining if you used the first two components
for subsequent analysis?
24)
Explain
some of the problems with using the mathematically derived components for image
analysis rather than the original bands. What are the benefits to this type of
approach?
25)
What are
the drawbacks? (HINT: What do you know about the original image bands that you
do not about the components?)